AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Molecular Docking Ppt3/29/2021
Essentially, molecular docking is a highly versatile tool that provides a nice push to an overly-slow industry, making the development of new drugs a bit less tedious and definitely more interesting in terms of understanding how proteins bind.In a world where data is being generated at a faster rate than we can process, bioinformatics is used to analyze massive amounts of data and make sense of it all.
 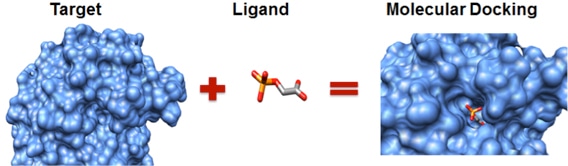 The expression in silico was first used in 1989, meaning performed on a computer or via computer simulation 1.The stages that follow the design of a new drug are both costly and time-consuming. The expression in silico was first used in 1989, meaning performed on a computer or via computer simulation 1.The stages that follow the design of a new drug are both costly and time-consuming.
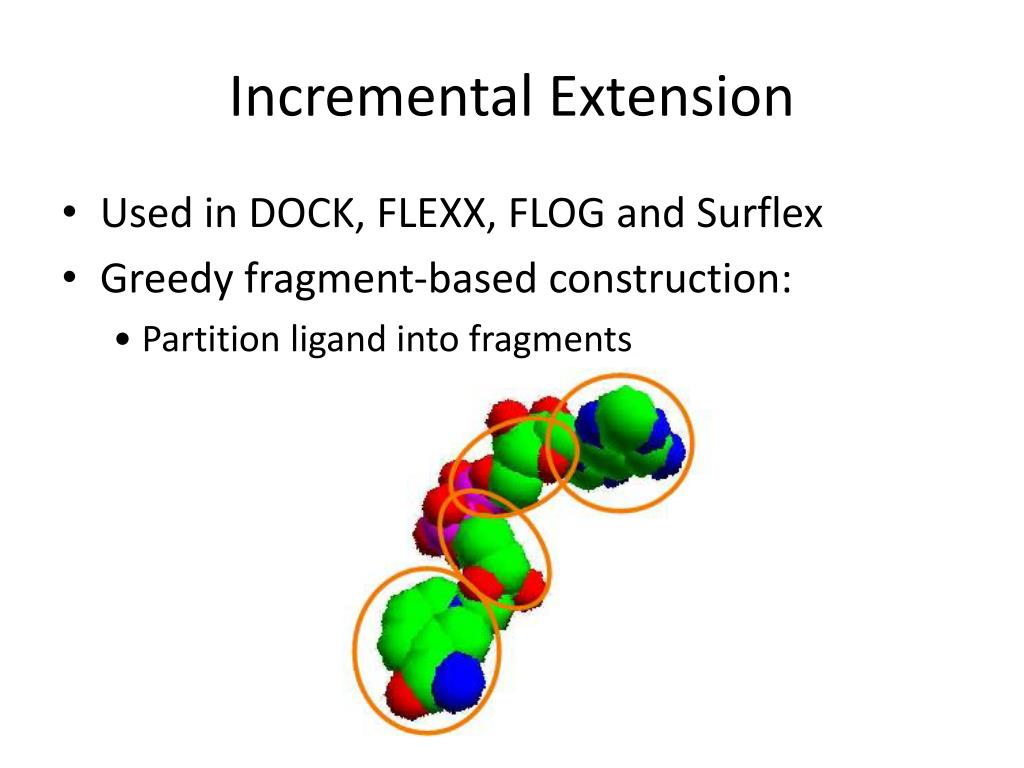
The entire process of drug development can take from 12 to 15 years and cost billions of dollars, but in silico studies have been seen to both speed up the discovery rate and reduce (although not eliminate) the need for expensive lab work. Typical stages of drug development from its discovery to commercialization 2 In silico research is usually performed during the drug discovery and optimization process comprising many computational methods such as homologycomparative modeling, molecular docking, virtual high-throughput screening, quantitative structure-activity relationship methods (QSAR), conformational analysis, the list goes on. Molecular docking a type of bioinformatic modeling, an essential tool in structural molecular biology and in drug design. The purpose of using this technique is to predict the most likely binding scenarios between a protein and a ligand, given their three-dimensional structures 4, 5. They are responsible for virtually every single biochemical process inside our bodies. Collectively, molecules that bind to a receptor are called ligands. By binding, ligands can either activate receptors (agonists) or deactivate them (antagonists). Computational power can be used to predict to a certain degree of accuracy where and how well a given molecule can attach itself to the receptor. However, while this technique might seem to be able to reveal potential drugs rather easily, in silico methods and simulations are definitely not a substitute for good ol laboratory assays Computational methods are simply not advanced and robust enough to simulate the exact interactions between large molecules. The binding interaction between a ligand and its target protein is usually a good gauge of its therapeutic activity 3. Molecular Docking Ppt Software To GenerateThe entire process is centered around using software to generate an enormous number of ligand-protein conformations, followed by calculations that predict which ones bind most strongly and are the most stable. 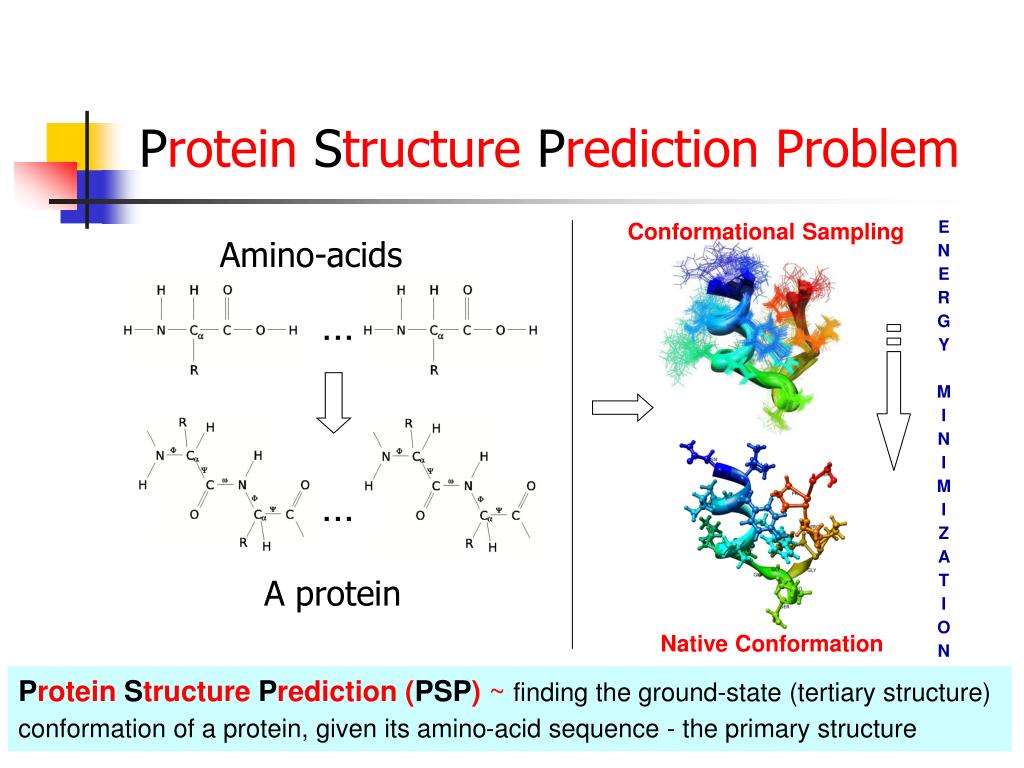
With the aid of sampling algorithms and certain assumptions, they are a key part of molecular docking as we can attempt to reproduce the binding event as accurately as possible 5, 6. Some extensions of these calculations include other relevant aspects such as hydrogen bonds or entropy. Molecular Docking Ppt Free Energy OfIt is important to take into account the solvent effects in these calculations, as it also plays a role in the free energy of the binding. The values generated by this statistical model are then used as coefficients to adjust the equation in general. These coefficients rely on the chosen dataset as well as the type of software used, making it common for differing results to be generated. Its advantage lies in its computational simplicity, with a drawback being some interactions might not be represented in the available database of crystal structures. Using machine learning tools, the operator can also choose a protein set to train the program with, producing different results.
0 Comments
Read More
Leave a Reply. |
- Home
- Services
- Team
- Contact
- Blog
- Hypermill file to solidworks
- Disc cover 3 supported printers epson printer 835
- Omnifocus keyboard shortcuts
- Fortnite aimbot map code
- Online photo exif metadata reader
- Keil 5 - tm4c1231c3pm
- Online rar extractor
- High school dreams best friends forever torrent
- Zynaptiq pitchmap ilok
- Corel draw x6 clip art download
- Icad
- Vocaloid editor 4 for cubase
- Home
- Services
- Team
- Contact
- Blog
- Hypermill file to solidworks
- Disc cover 3 supported printers epson printer 835
- Omnifocus keyboard shortcuts
- Fortnite aimbot map code
- Online photo exif metadata reader
- Keil 5 - tm4c1231c3pm
- Online rar extractor
- High school dreams best friends forever torrent
- Zynaptiq pitchmap ilok
- Corel draw x6 clip art download
- Icad
- Vocaloid editor 4 for cubase
- Home
- Services
- Team
- Contact
- Blog
- Hypermill file to solidworks
- Disc cover 3 supported printers epson printer 835
- Omnifocus keyboard shortcuts
- Fortnite aimbot map code
- Online photo exif metadata reader
- Keil 5 - tm4c1231c3pm
- Online rar extractor
- High school dreams best friends forever torrent
- Zynaptiq pitchmap ilok
- Corel draw x6 clip art download
- Icad
- Vocaloid editor 4 for cubase
 RSS Feed
RSS Feed
